Optimizing ASReview simulations: A generic multiprocessing solution for ‘light-data’ and ‘heavy-data’ users
Active learning can be used for optimizing and speeding up the screening phase of systematic reviews. Running simulation studies mimicking the screening process can be used to test the performance of different machine-learning models or to study the impact of different training data. This paper presents an architecture design with a multiprocessing computational strategy for running many such simulation studies in parallel, using the ASReview Makita workflow generator and Kubernetes software for deployment with cloud technologies. We provide a technical explanation of the proposed cloud architecture and its usage. In addition to that, we conducted 1140 simulations investigating the computational time using various numbers of CPUs and RAM settings. Our analysis demonstrates the degree to which simulations can be accelerated with multiprocessing computing usage. The parallel computation strategy and the architecture design that was developed in the present paper can contribute to future research with more optimal simulation time and, at the same time, ensure the safe completion of the needed processes.
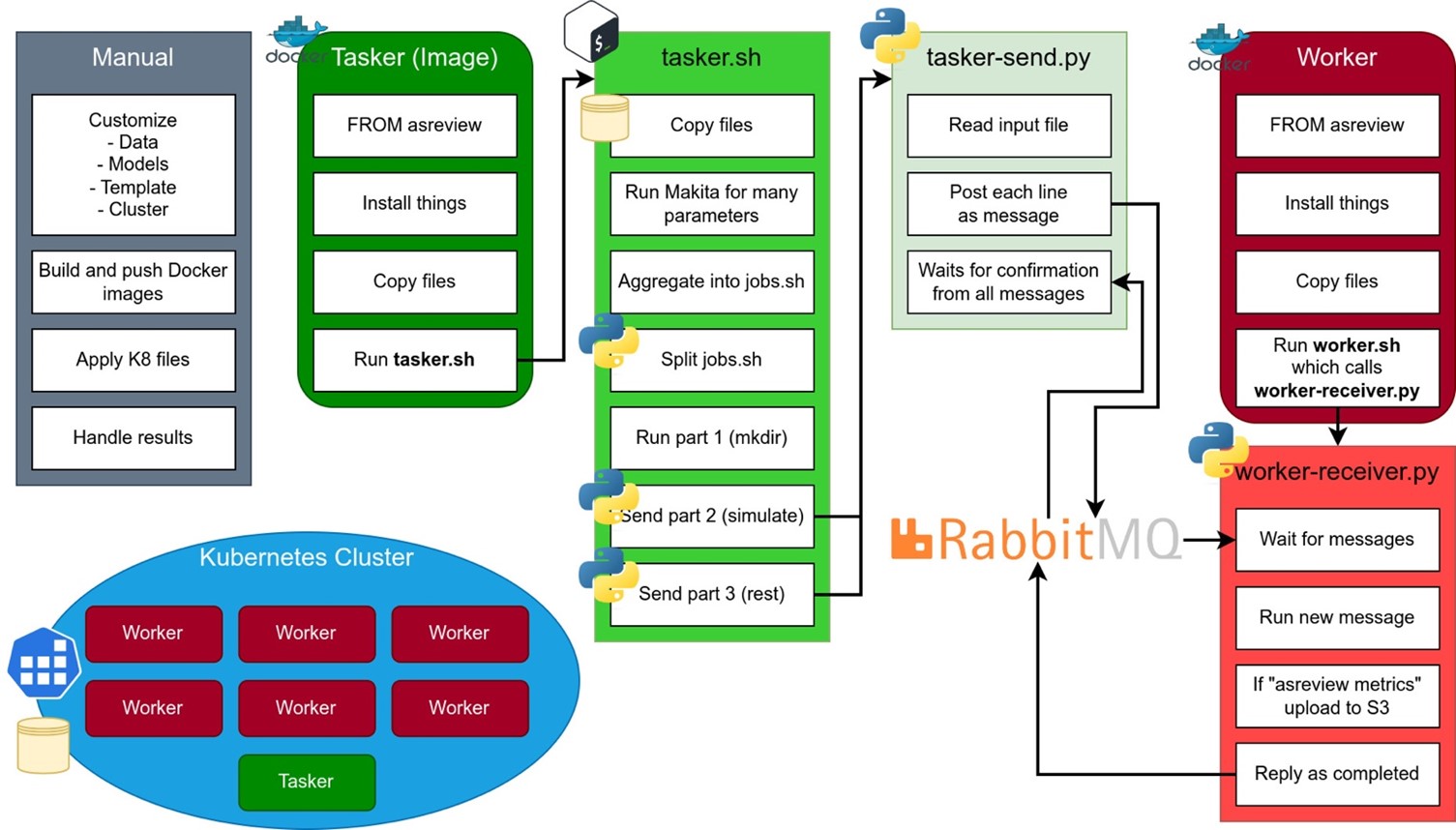
Kubernetes cluster implementation
Diagram of the ASReview parallelization design for cloud Kubernetes cluster implementation that describes the setup manual steps and the way two Docker components (Worker and Tasker) communicate using a ‘tasker.sh’ and ‘worker.sh’ bash scripts with the addition of RabbitMQ Message broker.
Documentation and Instructions
The presented architecture was provided with explicit documentation describing the steps necessary to run a large number of simulations via the cloud and local environments.
Scripts, data and output
Publication
Sergei RomanovAbel Soares SiqueiraJonathan de BruinJelle TeijemaLaura HofsteeRens van de Schoot; Optimizing ASReview simulations: A generic multiprocessing solution for ‘light-data’ and ‘heavy-data’ users. Data Intelligence 2024; DOI:10.1162/dint_a_00244
This work was supported by the Netherlands eScience Center under grant number ODISSEI.2022.023.